Developing a tool to easily store data and study details
Student: Dinara Issagaliyeva
Mentors: Jil Meier, Petra Ritter, Michael Schirner
Organization: INCF, Charite Universitätsmedizin Berlin
Introduction
The main problem that researchers face when exchanging data is a lack of consistency. They can spend hours
looking into the acquired data, trying to understand what each file represents. Since it's a time-consuming
task, Gorgolewski et al. (2016)[1] came together to create the
Brain Imaging Data Structure (BIDS)
format.
The specification covers a wide range of files having a unique set of rules for storing different kinds
of data. Several examples of the covered data sources are magnetic resonance imaging (MRI), electroencephalography (EEG),
and positron emission tomography (PET). However, it did not
cover simulation data conversion and validation.
Implementation
The GSoC'22 project was focused on building a user-friendly interface to convert computational data
(simulations) to a BIDS-compliant format and a validator to check the accuracy. The app was developed
following the principles of the BIDS Extension Proposal 034 (BEP034) by Dr. Michael Schirner and Prof.
Petra Ritter. The specification proposes a data structure schema for neural network computer models
that aims to be generically applicable to all kinds of neural network simulation software, mathematical
models, computational models, and data models, but with a focus on dynamic circuit models of brain activity.
Here is the list of final products created:
sim2bids
The project consists of an easy-to-use GUI that is built using the Holoviz Panel package. The approach I took to building the app was to minimize manual work as much as possible. Fortunately, my mentors gave me the green light to come up with ideas and implement what I had in my mind. The app includes several great features to ease researchers’ lives:
- First, it can run both locally and on a server, it's deployed on PyPI and can be accessed from anywhere.
-
Second, users do not need to go through a painful manual process of renaming files. They can pass the
files to the app, go to the
Preprocess Datatab, and select which files to rename into which standard naming schemes. This process is applied for every file having a unique extension. For example, if they have 100 subjects and there's a file called SC (structural connectivity), they can safely select the "weights" file from a drop-down menu. All files having "SC" will then be renamed. -
Third, users can change the converted JSON files directly in the app. For that, they need to navigate
to the
View Resultstab and select the file of their choice. There is an additional feature that changes the changed field in other files as well. Of course, the app will not change the critical parts that are unique for each file. -
Last, the app comes with an in-app user guide which can be found in the
User Guidetab. It explains how to use the app, explains any required files for renaming purposes, and more.
All the aforementioned functionalities are shown in the video below. The video shows the app running on an inline Jupyter notebook on the EBRAINS server.
Accepted files
The app supports a wide range of files, specifically: text (.txt), tab-separated values (.tsv), generic format (.dat), NumPy arrays (.npy), MATLAB files (.mat), HDF5 (.h5), and zip folders to extract network and coordinates files. It also supports three programming languages: Python, MATLAB, and R to extract model-specific parameters.
How to use the app
First, we need to download the package which can be easily done
using the following command:
!pip install sim2bids
Then, run the following commands in the Jupyter
notebook/Terminal/Python file:
import sim2bids
import panel as pn
pn.extension('tabulator', 'ace', 'jsoneditor', 'ipywidgets',
sizing_mode='stretch_width', notifications=True)
pn.serve(sim2bids.sim2bids.MainArea().view())
Output
Below you can see the output folder for simulation data conversion where each file is shown in its respective folders. For example, all coordinate files are stored in the global `coord` location. It is also important to note that these simulations were run on TVB v2.6. The app captures the source code to replicate the study.

comp_validator
One of the requirements for the project was to also create a BIDS validator for computational data
since it's not covered in the current form of the BIDS validator. The current BIDS validator app mainly
runs on the website where users can upload their folders directly. For our use case, it was important
to use Python-based implementation which can be run directly in the app or in the Jupyter notebook.
The custom BIDS validator for BEP034, comp_validator, has the same structure and rules as specified
in the previous
BIDS
validator. However, it also introduces naming conventions, data types, and accepted
files as specified in the
BEP034 proposal.
It outputs two different Markdown files covering errors and warnings.
BEP034
proposal. It output two different Markdown files covering
errors and warnings.
How to use the comp_validator app
First, we need to download the package which can be easily done
using the following command:
!pip install comp_validator
Then, run the following commands in the Jupyter
notebook/Terminal/Python file:
import comp_validator.comp_validator as validate
# specify path to the converted folder
validate.validate(PATH)
Output
Possible outputs of these commands are shown below:
Errors Markdown file
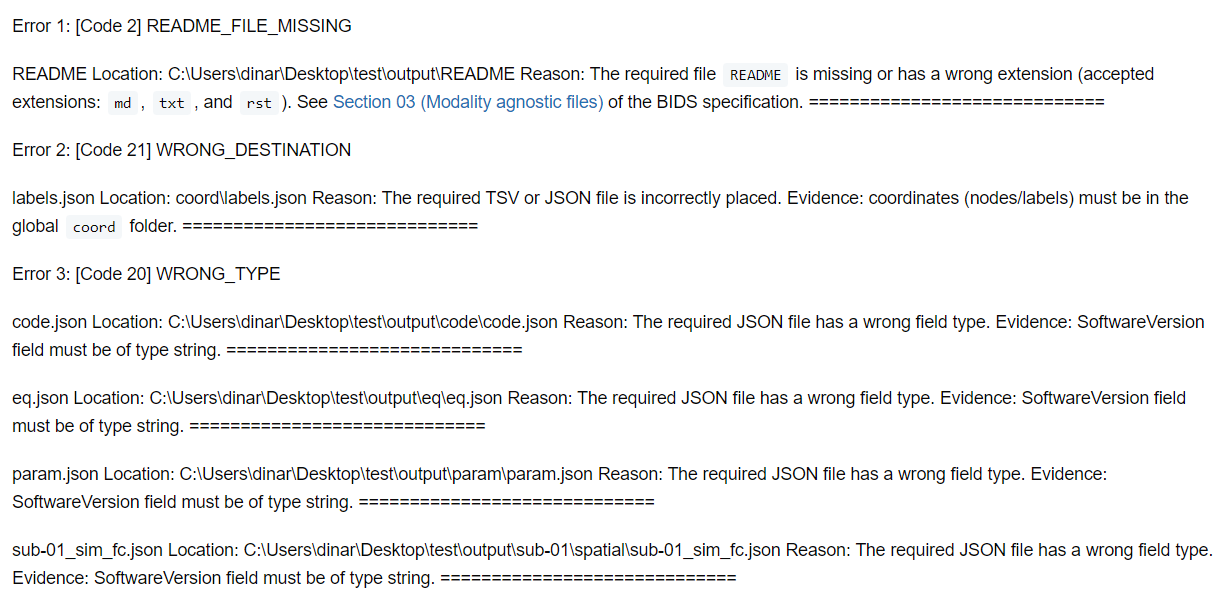
Warnings Markdown file

Achievements
The sim2bids app has been showcased so far in two large-scale educational events hosted by INCF and EBRAINS by Dr. Jil Meier in live-hands-on-coding sessions.
INCF Neuroinformatics Assembly
This was the first hands-on showcase where 50+ people joined online, and 500+ people downloaded the app and gave it a try!
Simulate with EBRAINS
This was the second hands-on showcase where 40+ people joined online. All the app's functionality was shown!
Additional links
There was a number of additional things created for the sim2bids showcase that you can access below:
Future Work
Even though the app can do quite a lot already, it didn't have a big exposure to different types of datasets. It's important
for its future to be maintained, adjusted, refactored, and perfected. It will also be beneficial to work on the app's promotion
so that other researchers know about the package.
A couple of things I'm planning to do after the GSoC completion:
- Integrate
comp_validatorto the app. Include a button to validate the dataset directly. Then, add an additional tab to show errors/warnings. - Refactor the code to decrease processing time.
- Write and publish a research paper about the project.
References
[1] Gorgolewski, K., Auer, T., Calhoun, V. et al. The brain imaging data structure, a format for organizing and describing outputs of neuroimaging experiments. Sci Data 3, 160044 (2016). https://doi.org/10.1038/sdata.2016.44